Hierarchical lineage tracing reveals diverse pathways of AML treatment resistance
Rachel Saxe, Hannah Stuart, Abigail Marshall, Fahiima Abdullahi, Zoë Chen, Francesco Emiliani, Aaron McKenna
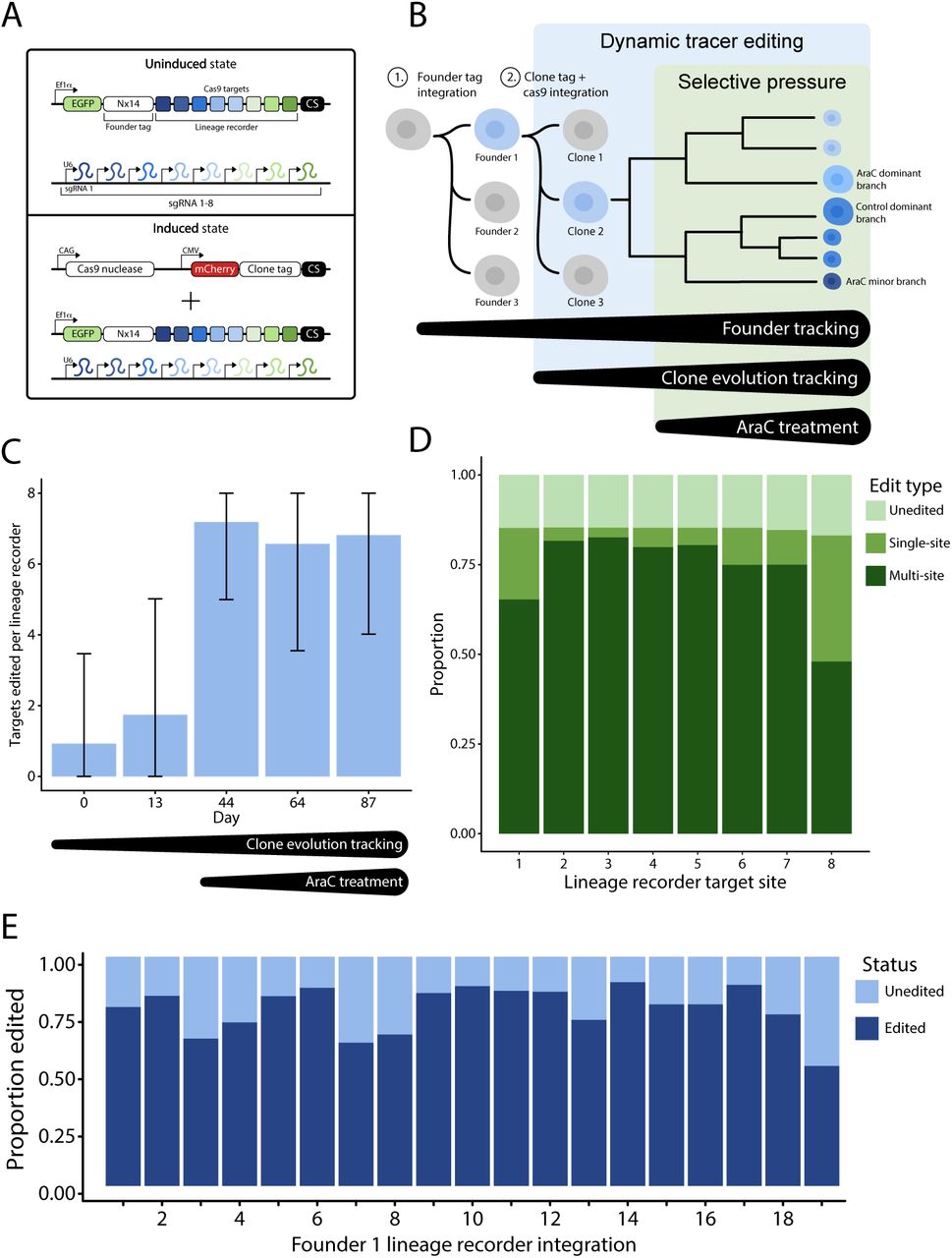
We developed FLARE (Following Lineage Adaptation and Resistance Evolution), a hierarchical, dynamic lineage tracing method for tracking cancer cell evolution under treatment pressure.
Using FLARE, we monitored acute myeloid leukemia (AML) cell lines exposed to Cytarabine (AraC), a standard AML therapy. We identified distinct cellular lineages in murine and human AML cell lines predisposed to AraC persistence and/or resistance via the upregulation of cell adhesion and motility pathways.
We additionally highlight the heritable expression of immunoproteasome 11S regulatory cap subunits as a potential mechanism aiding AML cell survival, proliferation, and immune escape in living organisms.
Our findings were validated against clinical data from the TARGET-AML cohort, revealing a broad spectrum of resistance signatures attributed to significant cell transcriptional changes. This represents the inaugural application of dynamic lineage tracing for investigating cancer treatment response mechanisms.